AQUATOX Frequently Asked Questions
Frequently Asked Questions by Category
- Capabilities
- Installation
- Source Code
- Example Study Files
- Biotic State Variables
- Initial Conditions
- Loadings
- Volume
- Sediments
- Parameters
- Libraries
- Ecotoxicology
- Waterbodies
- Models
- Output
- Metals
- Troubleshooting
Capabilities
Why use AQUATOX?
There are several reasons for choosing to use AQUATOX:
- “This is the first model that provides a reasonable interface for scientists to explore ecosystem level effects from multiple stressors over time” (peer review panel, 2008)
- Integrates fate and ecological effects
- nutrient and eutrophication effects
- fate and bioaccumulation of organics
- food web and ecotoxicological effects
- Predicts effects of multiple stressors
- nutrients, organic toxicants
- temperature, suspended sediment, flow
- Can be evaluative (with representative environments) or site-specific
- Peer reviewed by three independent panels and in several published model reviews (see Bibliography)
- Public-domain, Open Source code
How widely is AQUATOX being used?
The list server had 350 participants as of August 2013. However, it is difficult to know how widely the model is being used because we usually only hear from users when they have questions or problems! Known applications include:
- Criteria and maximum daily loads for nutrients and suspended sediments in rivers and streams in Minnesota, Idaho, Arkansas, Georgia, Florida, and Alabama
- Nutrient criteria for reservoirs in Arkansas, Oklahoma, and West Virginia
- Dissolved oxygen in a planned Turkish reservoir
- Management of nutrients in a Colorado reservoir
- Eutrophication of a lake in Finland
- Calibration and validation of biomass dynamics of periphyton and zoobenthos in experimental streams in France
- Ecology of a Greek river
- Fish dynamics in a North Carolina stream
- Fish ponds in Nigeria
- PCBs in shallow Dutch lakes
- PCBs in a Georgia reservoir
- Ecological assessment of pesticides in an Iowa reservoir
- Assessment of toxic organics from groundwater pollution of Danish stream
- Pesticide impacts in Dutch microcosms
- Estuarine ecosystems in South Africa
- A bay in Brazil
- Diversion of flood water from the Mississippi River into Lake Pontchartrain, Louisiana
- Risk assessment of chemicals for mosquito control in Long Island Sound
There is a bibliography listing publications that we are aware of that use or reference AQUATOX on the AQUATOX web site. An annotated bibliography that provides more details on these publications is also available.
Installation
What are the system requirements for AQUATOX?
Minimum:
- Windows 98, 2000, NT, or XP
- 1 GB RAM
- 100 MB free disk space
Recommended:
- Pentium PC, 2.0GHz or higher (or equivalent)
- Windows 2000, XP, or Vista, Windows 7 or 8
- 4 GB RAM
- 1 GB free disk space
I tried to install AQUATOX under Windows 7,8 (or Vista); why won’t it run?
Current Windows operating systems place restrictions on read/write access to certain directories as required by AQUATOX (“Program Files” is restricted for example). The InstallAware wizard for Version 3.1 suggests an appropriate default directory (the desktop) for installation; you may wish to change it. Please note:
- AQUATOX must be initially installed by a person with administrative privileges.
- If AQUATOX will exclusively be run using an “administrator” account, no additional changes need to be made.
- If AQUATOX will be run by a user with a “standard” account, full read and write privileges must be given to that user for the directory in which AQUATOX has been installed.
- An alternative to adding read and write privileges is to install AQUATOX in a directory where the user already has read and write privileges (the desktop generally works).
- See the Installation Guide.
Download the InstallAware wizard for Version 3.1
What is in an "APS" file?
An APS file is the basic unit in which AQUATOX simulations are loaded and saved. Each APS file (or “study”) contains the following:
- List of state variables and driving variables utilized and their loadings, "underlying parameters," and initial conditions;
- Site specific and remineralization parameters;
- Model setup information;
- Site constants and loadings for a simulation;
- Results from any simulation that may have previously been run, including "rates";
- External data that have been imported into the simulation for plotting against results;
- The graph library that has been produced for that simulation;
- The uncertainty mode or sensitivity mode setup including distributions chosen and correlation information.
The design criterion is to have a stand-alone file that contains everything except uncertainty and sensitivity analysis output. The intent is for a user to be able to archive the implementation (along with the current version of AQUATOX) to satisfy statutory and other requirements in case the analysis should be re-visited in the future.
How does an “APL” file differ?
An APL file links aquatic segments so that complex spatial relationships can be represented. Considerably more data are required for a linked-segment implementation, and often flow exchanges among segments will need to be developed offline, usually with a spreadsheet. Therefore, if possible, use an APS implementation, which is designed to be run with minimal data. However, if it is necessary to represent complex site-specific relationships then the APL implementation is available.
Source Code
How can I inspect the source code and compile it?
The open-source code can be downloaded without charge from the About Screen under Help by acknowledging the common public license. However, the AQUATOX model must be compiled with Delphi 2007 Professional, which must be purchased. If you buy the latest version of Delphi (XE3) you can obtain access to previous versions:
http://www.embarcadero.com/products/delphi/previous-versions Exit
Example Study Files
It seems like some of the studies are not realistic simulations; can you provide some guidance? I am especially interested in the nitrogen cycle.
First of all, you should heed the warning in the Technical Documentation that "the nitrogen cycle is represented with considerable uncertainty;" specifically, ammonia tends to be overestimated. Furthermore, the example studies come from many different applications, some of which were well calibrated and others that were only roughly calibrated.
For immediate reference, the best calibrated studies are:
- The Minnesota rivers (Crow Wing R. MN.aps, Rum R MN.aps, Blue Earth R.MN.aps, MN Rivers.als)
- The Lower Boise River, Idaho
- The Cahaba River, Alabama
- DeGray Reservoir, Arkansas
- Tenkiller Reservoir (Tenkiller Ferry Lake), Oklahoma
- Onondaga Lake, New York
Roughly calibrated studies include:
- Lake George New York, boundary conditions are questionable because only the southern part of the large lake is modeled, and the loadings were in part from extensive marshes.
- Coralville Lake Iowa, as calibrated, is dominated by loadings of phytoplankton in the inflow.
- Cheney Lake, Kansas, and Lake Jesup, Florida, were included as examples based entirely on the literature with only cursory evaluations.
- The Clear Lake, California, study was based very roughly on Clear Lake as an exercise in modeling herbicide use in an AQUATOX short course.
- Evers Reservoir, Florida, Skensved, Denmark, and Pyhajarvi, Finland, were set up and calibrated by other modelers, and are offered without support.
Biotic State Variables
Is there a way to enter additional species beyond the limits imposed by the wizard (6 algal species within groups except for “Other”, which has a limit of 2 species, and 2 invertebrate species within a guild)?
One possible workaround for the limit on the number of species within a group is to enter a new species in another group where we don’t have the maximum number of species allowed and just assign parameters as needed. The greens, diatoms, and “others” in the water column may be freely substituted with only one problem: the "percent diatom" and "percent greens" metrics calculated by the model will not be correct. Avoid substituting for blue-greens because they may float near the surface and fix nitrogen.
Guild designations for invertebrates and trophic-level designations for fish are used only to help represent possible relationships; these designations may be ignored and substitutions may be freely made within invertebrates and fish.
How would one model macrophytes in the littoral zone of a lake or reservoir?
Unless the lake is very shallow, the bathymetric relationship used for the whole lake would represent the littoral fringe as a small percentage; and the macrophyte biomass would be reduced by that percentage. (For more information, see the FracLittoral component of Technical Documentation equation (85) in which photosynthesis is calculated.) In other words, for a point model, macrophytes in the littoral zone are normalized by the volume of the entire lake to accurately represent their importance in the food web for the whole lake. If it is important to model a specific macrophyte-rich region, you should set up a linked littoral segment in either a lake or a reservoir.
Because macrophytes generally are assumed to get all their nutrients from the sediments, is there any way that I can model seagrasses that are taking nutrients from the water?
Yes, you can use Fontinalis, an aquatic moss, as the template for macrophytes that depend on nutrients in the water column. We will add genera such as Zostera to the library once they have been calibrated.
Initial Conditions
Biomass
What should I do if I don’t have good biomass estimates for initial conditions?
Because stream retention times are generally short, the model is not sensitive to initial conditions in streams, except for the sediments (including toxicants within the sediments). If you are modeling a stream the initial conditions in the water column are immaterial because upstream loadings will replace them fairly rapidly anyway. Lentic systems may be sensitive to initial conditions for periods less than three years. One option is to run the model in “spin-up” mode, which will save the biotic end conditions as initial conditions. You may also choose to spin up the sediments and nutrients, although this should be done with caution (to ensure that your model end results represent a steady state rather than reflecting a singular event).
Nutrients
The concentrations of nutrients in my estuarine site vary a lot spatially. So, for the initial conditions should I use a mean of all these values, or should I consider just a single point to represent the segment?
It depends on the type of variation: if the variation is spatially random then using an average of the data is probably best (and you can probably use the simple salt-balance version); if the variation is along the length of the salinity gradient, then using averages within linked segments is better. Also, you can spin up the nutrients from the initial conditions and run the model for a year to let the model respond to the boundary-condition forcings.
Toxicants
In the East Fork Poplar Creek TN PCBs.aps example study, how is the system loaded with PCB 1254? I have clicked on the chemical screen and looked in all the conditions and loadings boxes I can find, and they all show 0. Can you explain how AQUATOX is obtaining the PCBs?
This site represents legacy (prior) pollution. All the PCB loadings are sorbed to the detrital compartments in this study. For example, open the Refrac. Sed. Detritus loading screen and you will see on the right-hand side that there is 45 ug PCBs/kg dry associated with those organic sediments in the stream bed. The same concentration occurs in the Labile Sed. Detritus, and the Suspended and Dissolved Detritus has a declining concentration over the period of 1993-2001.
Why is there sometimes a nonzero value for a toxicant in the control simulation?
“Control” doesn’t always mean without toxicant. You might hold the chemical constant in both simulations and vary the detrital loading, for example. Calibration can be accelerated by turning off the differences and using both simulations to sequentially calibrate the model—comparing the newest with the previous, regardless of the designations “perturbed” and “control.”
Loadings
Flow
I am using the Manning option to import "Discharge of Water" data from an external (Excel) file of USGS flows. After running the model successfully and examining the output, it appears that the reach outflows (DischH2O, cu.m/d) are slightly changed from what I specified in the Excel input file. Why the difference?
AQUATOX does not use a daily time step but instead calculates results on a variable step-size basis. It uses Runge-Kutta methods to determine if the relative error is good enough to continue. Then, results are averaged on a daily basis for output.
The reason this is important is because the procedure will regularly solve time-steps that start in one day and end in another day. In this case, trapezoidal integration is used to determine the value for a given day, which is subject to some error, especially when compared to "stair-step" data. So even though the model is being driven by "known" discharge values, the calculated values would show some error.
Is there a way to represent a storm hydrograph with loadings that vary during the course of a day?
Release 3.1 can use hourly loadings for the following variables: all nutrients, organic matter, CO2, Oxygen, inorganic suspended sediments (sand/silt/clay), TSS, and Light. At this time, water flows themselves can only be represented daily, however.
Nutrients
In my simulation, total phosphorus (TP) increases during the winter time period from the last sample at the end of October 2006 to the next sample in April 2007 even though there is little or no inflow. Why would TP increase during the winter?
The model interpolates between the last loading in 2006 and the first value in 2007, giving a linear increase in concentrations during a period when there is little or no loss to photosynthesis. You need to add zero values for nutrient loadings corresponding to periods of zero inflow.
Detritus
We have an observation of organic matter in BOD units; how is that converted to detrital biomass?
For suspended and dissolved detritus, a conversion from BOD is provided that is a function of the refractory (“slow-reacting”) percentage associated with that inflow loading (or point-source or nonpoint-source loading as the case may be.) AQUATOX assumes that the 5-day BOD input from the user reflects the labile (“fast-reacting”) portion of organic matter and using the refractory percentage can therefore estimate the total organic matter loading. See the discussion in section 5.1 of the Technical Documentation for more information about this conversion (Equations 148b and 148c), and for information on how to parameterize the important “percent refractory” model input.
Temperature
I had 5 actual measured values for the temperature of the water over a year, so I used the time-varying temperature choice in the wizard and obtained a strange temperature curve.
Linear interpolation with only a few data points can introduce large errors in a simulation. The preferable approach would be to guess at an annual mean and range using your few observations and have the model generate a sinusoidal loading.
How does AQUATOX predict ice cover in lakes, streams, and estuaries?
Ice cover in freshwater systems, regardless of flow rate, is assumed to occur when the average water temperature drops below 3 degrees C. If a site is defined as an estuary, ice cover is assumed to occur at -1.8 degrees C based on a salinity of 35 parts per thousand.
When the temperature falls to 3 degrees C, why does the oxygen curve flat-line at the initial value (8.5 mg/L)?
In your simulation, DO would drop to 8.5 mg/L with ice cover if the loading is 8.5 mg/L; in the absence of reaeration, and since respiration exceeds photosynthesis, the loading maintains the DO value.
Enhanced Phytoplankton Retention
Should I always use the "Enhanced Phytoplankton and Zooplankton Retention/Washout" option in the Site screen? Can I still implement drift when I use this?
Volume
Why, although I select the "keep constant at initial level" option for water volume in the model set-up, do I get serious declines?
If you are modeling a stratified system then the “Water Vol.” output switches from representing the entire system in an unstratified state, to representing one of the individual vertical segments in a stratified state. This looks like serious declines in water volume at the onset of stratification. To get the total volume, when the system is stratified, you need to add the volumes of the epilimnion and hypolimnion segments; the total volume should remain constant. To toggle between segments when viewing output, right click on the graph and select “other segment.”
Sediments
Under what circumstances would one use each of the sediment options described in the Technical Documentation?
These options represent different aspects of suspended and deposited sediments:
- The submodel based on total suspended sediment (TSS) loadings is used with minimal data and very simple regression relationships for sediment deposition; it is appropriate for streams;
- The sand-silt-clay (SSC) submodel is similar to that for suspended and deposited sediments used in HSPF; it should be used with site-specific observations of inorganic sediments and of shear stress;
- A complex multiple-layer sediment submodel is intended to be used in simulating buried and exhumed pollutants; it can also be used in conjunction with the SSC submodel;
- The sediment diagenesis model proposed by Di Toro is intended to represent nutrient flux in anaerobic sediments; it is appropriate for many lakes and reservoirs.
For more information about each of these models, please see Chapter 6 of the Technical Documentation.
Parameters
What are the most sensitive parameters?
This depends in part on the individual studies. However, some parameters are practically global in their importance:
- For algae: light saturation, nutrient half-saturation constants, maximum photosynthesis, mortality;
- For invertebrates: maximum consumption rate, maximum respiration rate;
- For fish: mortality, bioenergetics parameters, and feeding preferences
- All species: optimal temperature and temperature range parameters.
Detritus is used to include non-living organic matter as well as its associated decomposers (bacteria and fungi). But refractory (suspended and sediment) detritus is transformed to labile detritus by colonization by bacteria, and the decomposition of labile detritus is caused by bacteria and fungi. What are the rates of colonization and decomposition of detritus? Is it possible to know the bacteria that are considered by the model?
The "colonization" and degradation rates are given in the Remineralization record, which you can open through the Site button. AQUATOX does not explicitly model the bacteria and fungi, but rather assumes that growth and activity will occur as an environmental response represented by general parameters. Those editable, general parameters are also given in the Remineralization record.
What is the significance of the “P:organics” and “N:organics” in the phytoplankton parameter screen?
AQUATOX tracks organic matter rather than separate nutrients, as many models do, and stoichiometry can be set by the user for each biotic state variable. Uptake or assimilation of nutrients is set for each plant in the respective plant parameter screen based on the ratios of nutrients to organic matter.
Is there a way to change the rate of nitrogen fixation for cyanobacteria?"
The rate of fixation cannot be changed explicitly. As the Technical Documentation states, there are two ways to trigger the process:
- If the concentration of inorganic nitrogen is less than half KN, the half-saturation for nitrogen, nitrogen limitation is turned off
- Nitrogen fixation can also be triggered by a threshold value of the ratio of inorganic nitrogen to inorganic phosphorus, which is selected and specified in the “Study Setup” screen. When the ratio falls below the threshold, nitrogen fixation is assumed to occur.
In the first case, there may be fixation in one group of cyanobacteria but not in others, depending on the KN values. In the second case, all cyanobacteria are affected.
When nitrogen fixation occurs, nitrogen is no longer assumed to be a limiting nutrient for relevant plants. Additionally, free nitrogen from outside of the model domain is added to the nitrate and nitrite pool when nitrogen fixing plants grow.
I could not figure out why, but it is not possible to change maximum consumption and respiration rates (g/g-d) for fish species (the input boxes are locked). Is that done on purpose?
Most fish are modeled using their mean weight to obtain allometric estimates of consumption and respiration rates. If you wish to override that approach, uncheck the “Use Allometric Equation to Calculate Maximum Consumption” and “Use Allometric Equations to Calculate Respiration” options near the bottom of the parameter screen.
Fish results for my site are given in terms of CPUE (catch per unit effort) and total fish (in number) for the sampling period. We also have relative weight data (actual weight of the fish/standard weight). I've looked for a good way for conversion to fish biomass (mg/l) but could not find one. Did you deal with a similar issue in the past?
Conversions from fish catch data to biomass are questionable, especially in a lake. Often, within the AQUATOX output screen, what we have done is to plot the observed data on the Y2 axis so that you can compare trends with biomass on the Y1 axis over periods of years. Otherwise, you have to be willing to assume an area covered by the sampling and the approximate mean weights, with unknown error.
Would you please explain a little more about the physical meaning in the food function?
The food actually available to a predator may be reduced in two ways:
where:
BMinpred is specific to the predator. However, the biomass available for predation is evaluated for each prey type.
Many animals will stop feeding to avoid wasting energy searching for a scarce resource. They will concentrate on the abundant food. This function partially represents prey switching. Fishermen are quite aware of this phenomenon. If there is a mayfly emergence, the smart fly fisherman will switch to a fly that looks like an adult mayfly.
I am having a tough time maintaining the biomass of my fish - they lack food. Is this because my invertebrates are benthic and most of my fish are invertivores in the water column?
There are several considerations. First, check the BMin to make sure it isn’t set too high. Then look at the trophic matrix to make sure that there is a feeding preference for the zoobenthos; unfortunately, we can’t model external insect food sources at this time. If you are modeling a stream, perhaps a work-around for terrestrial insects would be to have a labile particulate load with a corresponding preference among the fish; the problem is that this could exert an additional oxygen demand.
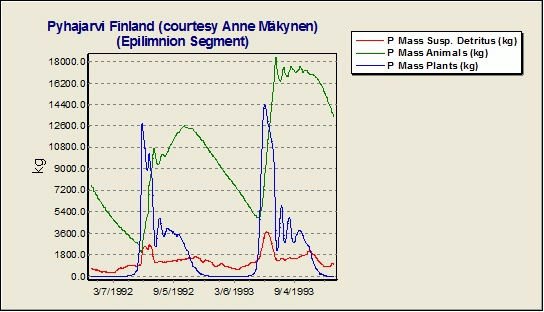
Why is there is such a large difference between the observed and simulated phosphorus concentrations in the hypolimnion in the Lake Pyhajarvi Finland example study?
One should be careful to implement fishing pressure in the model. Yearly fishing is removing almost 25% of the phosphorus load from Lake Pyhajarvi Finland; this corresponds perfectly with the difference between observed and simulated concentrations. AQUATOX provides output of total masses of nutrients tied up in various ecosystem components, including fish. The Lake Pyhajarvi example without removal of P by fishing is shown below.
Libraries
How do I update the libraries with my records when an update to AQUATOX is released?
The easiest way is to save your updated records to the new libraries. The program should update the new records automatically, and incorporate those into the library. Saving the chemical records should also save the associated ecotoxicology records automatically. Furthermore, in Release 3.1 you can save all the records in your study to a separate file, which is especially helpful in summarizing the parameters for documentation and publication. If you send us your studies we will try to include new records in the libraries that are distributed with successive releases. However, changing existing records in the default libraries should not be taken lightly, as these data should be retained as reference “starting points” for calibration.
How can I create, save, and use empty versions of the AQUATOX databases/libraries that I can then populate with my own sets of parameters?
Unfortunately the Paradox format has largely been abandoned, but we haven't had the opportunity to change the AQUATOX database structure yet.
However, there are some workarounds:
- To work with the Paradox data, you will need both the DB and PX files. If you are working with an "*.ADB" file or "*.PDB" file, you should rename that to a "*.DB" file before trying to read with external software. (These DB files are often readable by Excel, but you cannot usually save your changes back to DB without causing problems.)
- To create a new blank library, probably the best procedure is to copy the DB and PX files to an alternative file name (Blank_Animal.ADB and Blank_Animal.PX) and then delete all entries through the AQUATOX interface.
- To work collaboratively with these databases, it may be best to export from "grid mode" to Excel and then comment the cells as you are changing them before sharing them with your team . However, this will require entering all of those changes back into AQUATOX by hand at the end of the process.
How can I add new entries to the library (e.g. a species that is not currently in the database).
You can use the “New” button within the library interface to create a database entry that has all blank parameters. You can then name that entry appropriately and enter values for all parameters.
However, it often is easier to use a surrogate “most similar” species and then modify the existing set of parameters. The easiest way to do this is to add the “most similar” species into an AQUATOX simulation, change the name of the species and the relevant parameters, and then save that new species (with a new name) back into the AQUATOX Library. This same procedure will work with database entries for animals, plants, sites, chemicals, and remineralization records.
Ecotoxicology
The only information I have on the fate of a particular chemical is its half-life; how can I enter this?
Convert it to a rate (0.693/half-life) and enter it as the uncatalyzed hydrolysis rate.
Does the model deal with the fact that a user can enter different exposure durations for species for one compound? For example in the default database, i.e. for 2,4-D Acid, there are lots of animals, where the LC50 is given. Some of the tests were for 48 hours, while others were for 96.
When calculating toxicant effects based on internal toxicity (the model default), AQUATOX explicitly accounts for the differing exposure times when computing an infinite LC50. Because time of exposure is accounted for in model calculations, the model will predict 50% mortality after 96 hours of exposure when working with a 96-hour LC50. See the section on “Internal Calculations” in Technical Documentation section 9.1. When calculating toxicant effects based on external toxicity, effects are assumed to occur instantaneously.
I'm trying to simulate the effect of a chemical in my system. But every time I run the simulation with an organism (plant or invertebrate) it gives me a Fatal Parameterization Error: that the organism uses a toxicity record that is not found in the chemical toxicity data. How can I get this to work?
If you double click on the chemical in the state variable list and then click on the "toxicity data" button you will see the available toxicity records for animals and plants. In the underlying data for each animal and plant, the "toxicity record" field must match one of these records in the chemical.
So you have two options. Either populate your chemical's toxicity screen with more records (using available data or the EPA ICE database of least-square regression models available through the "Interspecies Toxicity Correlation Models" button), or set up the animals and plants in your study to match the existing records. (You can assign all existing animal and plant toxicity records simultaneously using the "edit all" button in the upper right corner of the organism screen.)
Don't forget about the context-sensitive help available on each screen, especially for the complicated Toxicity Correlation window.
Waterbodies
Maximum Size
What is the maximum size for a waterbody before it should be represented by linked segments?
It depends on the modeling goal. If the model is being used to evaluate potential bioaccumulation and toxic effects of a chemical for purposes of registration, then a point model or stratified 1-D model may be sufficient use of broad, representative environments (such as in the Galveston Bay example). However, if the goal is analysis of site-specific pollution, then linked segments probably should be used for all but the simplest case.
The Technical Documentation states:
According to Ford and Thornton (1979), a one-dimensional model is appropriate for reservoirs that are between 0.5 and 10 km in length; if larger, then a two-dimensional model disaggregated along the long axis is indicated. The one-dimensional assumption is also appropriate for many lakes (Stefan and Fang, 1994). Similarly, one can consider a single reach or stretch of river at a time.
Streams
I have been running 6 stream segments in cascade mode. I just switched over to feedback mode for the links, assuming a 5% fish migration to adjacent segments once a year. I was surprised at how different the result was. What else changes in feedback mode? Asked another way - if I were to change the links to feedback but not specify any migration, should I expect any difference, and if so why?
In cascade mode the loadings between segments are daily, whereas in feedback mode they occur at whatever the minimum step-size dictates since all equations are solved simultaneously. This is especially important for diel simulations of DO, where results will get passed at hourly or smaller time steps. I think you will find very little difference between the cascade and feedback runs with no migration.
Are the habitats (run, riffle, and pool) modeled separately?
No, the model just calculates the velocity for each of these separately and determines the organism's response to velocity depending on where it is located.
Ponds
How can I adapt AQUATOX to an aquaculture system?
If you are referring to fish ponds, I suggest you use Farm Pond MO.aps or Farm Pond MO Esfenvalerate.aps, which are distributed with the model. The latter study serves as the example for the Simple Tutorial in the User’s Manual and Help screens. The simulated fish species should be those that you would stock in your system. After you have worked through the tutorial you should feel free to experiment. The pond study runs very fast, so you can try many parameter combinations as you calibrate with your data.
Mesocosms
What types of experimental waterbodies (mesocosms) can AQUATOX simulate?
In keeping with its original development as an evaluative model, AQUATOX can represent the full range of mesocosms. These include small ponds, experimental streams, and enclosures in ponds and lakes. In the latter (sometimes called “limnocorrals”), the area of the enclosure wall is explicit, and is set in the site screen.
Marine
Can AQUATOX be used for marine-related applications, apart from the estuarine option?
The only reason to hesitate to use it for open marine applications would be because of the difficulty in defining the flow field. However, that could be solved by using the output from a hydrodynamic model with a linked-segment implementation of AQUATOX. For pH and CO2 calculations, linkage to the CO2Sys model is now available, as discussed in section 5.6 of the Technical Documentation.
I have a question about how to use my volume and area values in my study. My problem is that in my bay the mangrove area is significant, and therefore I must be very careful to choose the sea level reference, since any difference will result in large variations in the area. Should I consider the level of water as being of low tide, mid-level or high tide?
Because AQUATOX does not represent wetlands, you should use low tide as the reference.
I want to model a lagoon that is long and skinny, with a very narrow outlet at the top. Do you think that AQUATOX represents dynamics for something like this very well? The wide vs. narrow portions of the lagoon are rather different. Would I want to use the segmented version?
The standard estuarine model has two layers: brackish and salty. The salt balance approach does not consider the width of the opening but rather is driven by the observed time-series of salinity in the two layers. Presumably, the narrow opening would affect the salinity. (In contrast, one user was modeling a bay that had such a wide mouth that it was well mixed, and the modeler had to use data from inside the mouth.) Keep in mind that any implementation of an estuary is going to be a simplification. In your case I would use a simple segmented model with salinity as a driving variable.
Will the model allow simulation of a conservative tracer or constituent that could be used as a surrogate for salinity to develop model projections for flow among segments?
AQUATOX does not have a conservative tracer. However, you could fashion your own tracer by changing a toxicant to what is essentially an inert dissolved compound. Set all degradation rates to zero, including hydrolysis and photolysis. Ignore the activation temperature. Set KOW to a very small value in order to minimize sorption. Set the Henry’s Law constant to 0 to turn off volatilization. Those settings should result in a "conservative" compound.
Link to Watershed Models
AQUATOX now has two available linkages to the HSPF watershed model. Will the updated linkage to HSPF replace the current linkage, or is it a new alternative?
This is a new alternative, triggered from the AQUATOX menu and reading WDM files directly, rather than triggered through linkage from within BASINS using WinHSPF.
Can the current version link to BASINS as well as stand-alone HSPF?
The new one could link to BASINS as all HSPF simulations read and write their data from WDM files.
Does the new linkage have additional capabilities?
The old linkage required users to use WinHSPF to set up AQUATOX-friendly outputs—the outputs included masses passing over boundary conditions, inflow volumes of water, and all had AQUATOX-friendly units and time-steps. The new linkage is far more flexible:
- It doesn't assume the units or time-steps used in HSPF and checks those units to make sure that they are as expected. If units are not specified in the WDM file the log-file alerts the user that it assumes they are using a certain type of unit (general HSPF defaults). If it finds unexpected units it raises an error.
- It reads in-stream HSPF concentrations rather than requiring the HSPF simulation to specify information about boundary conditions.
- It converts units from the often-used HSPF British units into AQUATOX units in several cases (temperature, depth, water flows)
- It uses TOC as the default organic matter input if it's available from HSPF, otherwise it reverts to BOD.
- It is more flexible with regards to phosphate linkage. The user can select which HSPF compartments they wish to link to which AQUATOX phosphate compartments (i.e. TSP or TP).
Output
How can one read the output files produced by uncertainty analysis?
AQUATOX outputs Paradox databases. The databases are sometimes viewable in Excel, especially older versions. There are free viewers available: https://en.wikipedia.org/wiki/Paradox_(database) Exit
Because of limitations in the Paradox database, tracked results are split over many DB files. These can be selected and viewed in AQUATOX. We recommend using the "save each iteration to CSV" option and performing data analysis in Excel using the CSV file rather than relying on the limited db outputs.
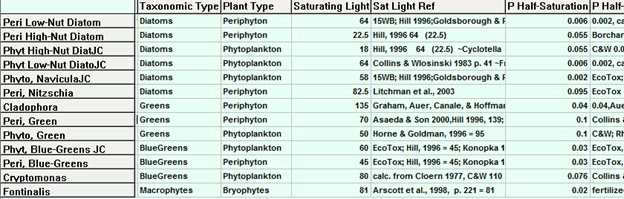
I want to show biological parameter data in tabular form for my paper. Is there any handy way to get data about fish, invertebrates and phytoplankton into Excel or Word from the parameter sheets in AQUATOX?
Release 3 allows the parameter records for those groups (plants or animals) to be displayed in GridMode (shown below). The user then has the option of saving the entire set to an Excel file, which can be edited for inclusion in the paper.
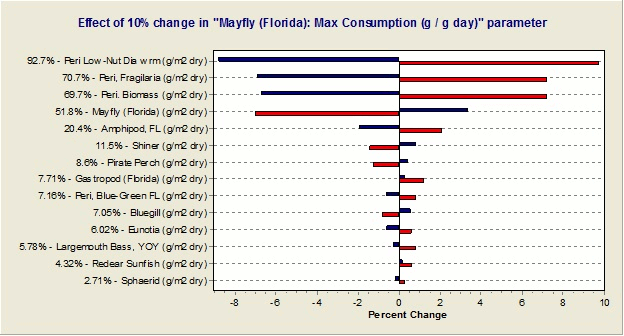
Would it be possible to determine the sensitivity of the simulations to each parameter?
Yes, one can implement a "reverse tornado" or "effects” diagram that shows the effects of each parameter change on the overall simulation, as shown below. (The vertical line at the middle of the diagram represents the deterministic model result. Red lines represent model results when the given parameter is reduced by the user-input percentage while blue lines represent a positive change in the parameter.)
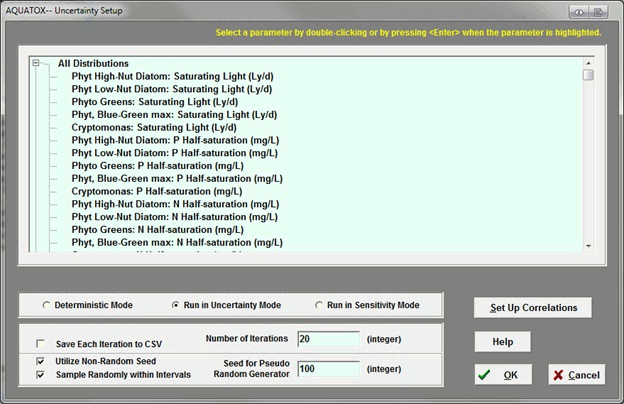
Can AQUATOX account for correlated variables, such as N and P, in uncertainty analysis?
Yes. There is a button on the right side of the uncertainty setup screen to allow the user to specify correlations.
When data are presented as percentages, they must always be relative to something. What is volatilization relative to?
The percentage is expressed in terms of percentage of the mass of chemical at that given time. So if you had a value of 5% that would mean that 5% of the chemical volatilized during that day. For example, with 100 ug/L of chemical, if 5 ug/L volatilizes, the rate is expressed as 5% volatilization. If you want the total mass of chemical that has volatilized since the start of the simulation, look at "T1 Volatil (kg)."
Why is there sometimes a difference in variables between the control and perturbed output?
If the simulation changes between the control and the perturbed simulations (for example, algal type or number of toxicants), the control and perturbed graphs may have different variables plotted.
Why don’t refractory and labile sediment detritus show up in the detrital output?
When running the “classic” AQUATOX sediment bed model, these categories appear as “R detr sed” or “refractory (slow-reacting) detritus in the sediment bed,” and “L detr sed” or “labile (fast-reacting) detritus in the sediment bed.”
If you are using the sediment diagenesis submodel, the compartments are expressed as particulate organic carbon (POC) occurring in the second layer of the sediment bed “L2 POC.” As POC is broken into three classes of reactivity, labile corresponds with G1 or the fastest reacting POC class, and refractory corresponds with G2 and G3. Nutrients associated with this organic matter POP and PON are also individually tracked in the sediment diagenesis submodel.
Which one represents the total sorption of chemical in the detritus sediment: T1 sediment, T1R detr sed(ppb), or T1L detr sed(ppb)?
The total sorption is best represented by the T1 Mass Bottom Sed (kg). You can add the T1 Mass Susp. Detritus (kg), but it is usually insignificant. The ppb results are concentrations.
Metals
Is it possible to add mercury to the chemical list? Are there some limitations in the model because mercury is not an organic chemical?
Methyl Hg is not lipophilic, so its bioaccumulation cannot be simulated using the default functions in AQUATOX. Years ago we had Hg in the model. However, we found it impossible to formulate a general model for methylation, and eventually we stripped out all the Hg code.
Has anyone tried to set AQUATOX up to model fate of a metal in the aquatic environment? I know it does not generally support such analysis at this time but thought someone may have played around with it or perhaps it is planned for the near future.
Recently a research group in Luxemburg has taken on the task of adding metals. They anticipate a fairly lengthy process supported by extensive verification and validation data.
Also, AQUATOX has been used to model the effects of copper sulfate as an algicide. Although the model does not represent the fate of metals, an equilibrium model such as MINTEQ can be used for that purpose. The user set up AQUATOX for external toxicity (see Setup screen) and drove it with observed copper loadings. The results were reasonable and tracked observed algal biomass in a reservoir dosed with copper fairly well.
Troubleshooting
Simulation Stops
Why does my new study get an error when I start to run it?
There are many possible reasons; if the error is immediate then a required boundary condition may be zero. For example, in a recent application the depth of the thermocline was set to 0. Errors that appear after the simulation has run for a while may be due to mistakes such as having an inflow that is consistently less than the outflow (AQUATOX will step through short-term dry conditions, but there is a limit to how long it will do so).
I get part way through a simulation that includes sediment diagenesis and the run “freezes.” What can I do?
First, you should choose the new option to simulate the aerobic layer as steady-state. This has only minor effects on model results and yet can prevent a diagenesis simulation from freezing up due to stiff differential equations. (When the steady-state model is not utilized, the state variables in the very-thin top sediment layer are solved using differential equations.) Second, if the model still freezes, increase the Relative Error in the Setup screen by a small increment. Also, you should always choose, in Setup, to have the model Show integration information; it will show the last rate that caused an error overrun in the simulation.
Simulation is Very Slow
I would like to know what the "percentage of maximum stepsize" really means when I run the model. I noticed that some changes that I make in initial conditions (especially in the water volume data) make big differences in the maximum stepsize, causing the model to be really slow or fast.
AQUATOX uses a variable-stepsize differential equations solver. This means that when model predictions have strong and frequent discontinuities (e.g. flashy streams), the equation solver reacts to the high relative error, reduces the stepsize, and tries to solve the equations again. If and when the response becomes less abrupt the model speeds up again. So that the user has a sense of how the solver is doing, AQUATOX can output the current step-size as a fraction of the maximum stepsize. The maximum stepsize is 1 day for a simulation with "daily simulation" selected in the setup window and 1 hour for a simulation with "hourly simulation" selected.
Changing the water volume can have a big effect on how rapidly inflow loadings change concentrations within a system. Therefore, it is not surprising that making a change in initial-condition volume could have this effect on the speed of the differential equations solver. If simulations get too slow, the fixed step-size option may be chosen.
Why would my simulation for a 10-year period take more than 12 hours?
That is an extreme result, even for a complex linked-segment run. There are several possibilities and corresponding remedies—all of which are good modeling practice:
- Sediment diagenesis may be the culprit; choose the new option to simulate the aerobic layer as steady-state. That will speed up the simulation considerably with very little additional error.
- The simulation may be set up so that extrapolation of time series slows down the run; if the model has to extrapolate a loading because it occurs outside the simulation period, the run time can be increased by as much as 15-fold in our experience. One work-around would be to generate the extrapolation offline and then import the time series into AQUATOX.
- If you use feedback mode, the stepsize for solving all differential equations will be equal to the smallest stepsize in any one of the segments. If you can, use cascade mode instead.
- Unless you really need hourly results, use a daily reporting step. It should speed up the run by at least a factor of 4 and will require much less memory.
Computer Goes into Inactive State
How is the computer prevented from going into sleep or hibernate mode during a long simulation, such as with a lengthy sensitivity analysis?
There are two processes that are utilized within the AQUATOX code to prevent system standby or hibernate in the event of a very long simulation:
- The software is queried by Windows in the event of system standby/hibernate/shut-down and asked whether to allow this. AQUATOX is set up to say "no" if it checks and a simulation is running.
- AQUATOX can declare the simulation-running threads as "critical" to the system so that the system won't go to sleep when a simulation is running as it is not officially "idle."
However, in some closely-managed computer systems, you may need to have your system administrator prevent these processes from being overridden.
