ToxCast Data Generation: ToxCast Assays
ToxCast uses automated chemical screening technologies, called high-throughput screening assays, to expose living cells or biological macromolecules such as proteins to chemicals. The cells or isolated proteins are then screened for changes in biological activity that may suggest potential health hazards.
- Cell-based in vitro assays use intact cells to measure cellular changes in response to exposure with the test substances. Example cells include human primary cells and cell lines and rat primary liver cells. An example is ACEA_T47D.
- Biochemical in vitro assays measure the activity of a biological macromolecule, either a purified protein or a nucleic acid. An example is NVS_ADME_hCYP19A1.
ToxCast uses more than 700 high-throughput assays that cover a range of high-level cell responses and approximately 300 signaling pathways.
Click links below for an overview of ToxCast assays and the platform sources, descriptions, and protocols to test chemical activity in high-throughput screening.
ToxCast Assays
- Overview
- Platform Sources, Descriptions, and Protocols
- Descriptions
- Information and Quality Statistics
Assay Process Overview
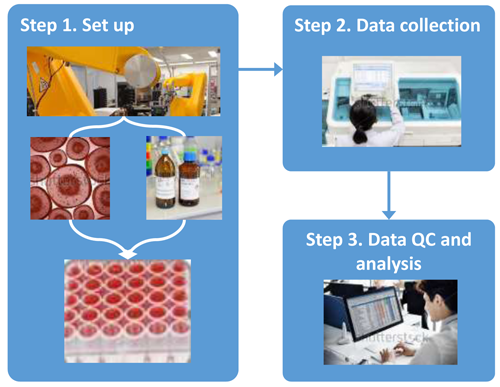 Step 1. High-throughput technology perform repetitive motions of loading the test chemicals, positive/negative control chemicals, dilution solvents, reagents, and cells (if applicable) into plates containing numerous wells (e.g., a 384-well plate).
Step 1. High-throughput technology perform repetitive motions of loading the test chemicals, positive/negative control chemicals, dilution solvents, reagents, and cells (if applicable) into plates containing numerous wells (e.g., a 384-well plate).
Step 2. The well plates are incubated for different time durations depending on the assay. Detection technologies translate the biological or physical process into a signal (e.g., fluorescence). The signals are measured and compiled into databases.
Step 3. Results are quality control checked, normalized, analyzed, and formatted for release. For more information visit Results Processing and Analysis.
